Downregulation of LPAR1 Promotes Invasive Behavior in Papillary Thyroid Carcinoma Cells
Abstract
Background:
Lysophosphatidic acid receptor 1 (LPAR1) has been identified as a biomarker in various cancer types. However, its biological function in papillary thyroid carcinoma (PTC) remains unknown.
Methods:
LPAR1 was identified as a key regulator of epithelial-mesenchymal transition (EMT) in PTC cells through bioinformatics analysis of TCGA and GEO datasets. PPI analysis and correlation with immune infiltrates were also conducted. LPAR1 expression was evaluated using Gepia2 and GTEx, and miRNA target gene prediction was done with multiMiR. To assess the expression of LPAR1, we extracted total RNA from both the BCPAP cell line and the normal human thyroid epithelial cell line Nthy-ori 3-1. The levels of LPAR1 expression were then measured using quantitative real-time polymerase chain reaction (qRT-PCR) in the BCPAP cell line, with a comparison to the Nthy-ori 3-1 cell line.
Results:
1081 genes were upregulated, and 544 were downregulated compared to normal tissue. LPAR1 was identified as a key candidate by analyzing the TCGA and GEO datasets. PPI data analysis showed interactions with metastasis-related proteins. Functional enrichment analysis indicated involvement in signaling pathways like phospholipase D and actin cytoskeleton regulation. LPAR1 expression correlated positively with immune infiltrates such as CD4+ T cells, macrophages, neutrophils, and myeloid dendritic cells but negatively with B cells. Additionally, miR-221-5p was predicted to target LPAR1 in PTC. Furthermore, our experimental data demonstrated that LPAR1 was under-expressed in the PTC cell line compared to the nonmalignant one (P < .01).
Conclusion:
LPAR1 suppresses metastasis and is linked to EMT, as evidenced by the decreased LPAR1 expression and increased miR-221-5p in PTC. This suggests its potential as a biomarker for diagnosis and prognosis and as a therapeutic target for EMT.
Introduction
The most common endocrine malignancy and the fifth most common cancer type worldwide is thyroid cancer, which is increasing (>5% per year).1 Thyroid carcinoma has several factors contributing to its onset and progression, although the exact etiology is still under investigation. These factors include ionizing radiation exposure, environmental pollution, smoking, hormonal imbalances, metabolic diseases such as obesity, and gender.2,3 Histopathology has identified several subtypes of thyroid carcinoma, with papillary thyroid carcinoma (PTC) accounting for 85% to 90% of all thyroid carcinomas.4,5 Other thyroid cancer subtypes include follicular thyroid carcinoma, anaplastic thyroid carcinoma, and medullary thyroid carcinoma.6 Despite a favorable long-term prognosis, the relatively high recurrence rate of PTC presents a significant challenge, especially when it comes to diagnosing and treating metastatic recurrence in advanced cases.7,8 In addition, overdiagnosis and overtreatment are common, especially in indolent cases. Identifying and treating indolent thyroid cancer cases can expose patients to unnecessary risks.9 Molecular-based studies can aid in diagnosing and treating PTC by providing insight into its underlying mechanisms.10,11 Consequently, there is an urgent and compelling need to develop novel and highly sensitive biomarkers that accurately identify specific PTC patients. This will enable personalized treatment strategies.
Lysophosphatidic acid receptor 1 (LPAR1) is a member of the G protein-coupled receptor family that binds to lysophosphatidic acid (LPA), a small phospholipid commonly found in serum, ascitic effusions, and inflammatory fluids.12 LPAR1 is pivotal to numerous biological functions, including chemotaxis, cell proliferation, differentiation, platelet aggregation, and tumor progression.13-16 Multiple cancer cell lines and primary tumors exhibit abnormal LPAR1 expression.17 LPAR1 has been identified as a prognostic biomarker for different types of cancer in several studies. They emphasize its crucial involvement in prostate, gastric, and pancreatic cancers.12,18-20 However, the specific functions and mechanisms of LPAR1 in tumor hallmarks, such as epithelial-mesenchymal transition (EMT), remain largely unknown. EMT, a process in which epithelial cells acquire mesenchymal characteristics, occurs in various biological processes. EMT plays a significant role in promoting tumor malignancy by enhancing invasiveness and metastatic potential.21
However, the biological role of LPAR1 in PTC remains unknown. In this study, we sought to investigate the expression of LPAR1 in PTC patients undergoing EMT by analyzing data from The Cancer Genome Atlas (TCGA) and the National Center for Biotechnology Information Gene Expression Omnibus (GEO) databases. Through comprehensive multidimensional analysis, we explored the genomic functional networks associated with LPAR1 in PTC and delved into its potential role in EMT. Our results shed light on LPAR1 as a promising and novel target for diagnosing and treating PTC. This highlights its significance in furthering our understanding of the disease.
Materia and Methods
RNA sequencing data preprocessing
On August 12, 2023, we accessed the GDC data portal (https://portal.gdc.cancer.gov/) to obtain RNA-Seq data and clinical information related to the TCGA thyroid carcinoma project (project ID = TCGA-THCA). The dataset consisted of 505 primary tumors and 59 adjacent non-tumor tissues for mRNA analysis. We removed duplicate samples and those without complete clinical-TNM stage data, resulting in a final count of 279 samples (203 females and 76 males) for primary tumors and 59 samples (41 females and 17 males) for adjacent non-tumor tissues.
Expression analysis of RNAseq data related to the THCA project
We used the DESeq2 package in R to conduct mRNA expression analysis, specifically comparing PTC samples with normal ones. Our analysis assessed differential expression between primary tumor and normal tissue samples. Genes were considered differentially expressed if they satisfied the criteria of a False Discovery Rate (FDR) of less than 0.01 and an absolute log2 Fold Change (logFC) greater than 1.
Identification of proteins related to epithelial-to-mesenchymal transition (EMT)
To investigate the role of proteins in EMT, we analyzed the GSE6004 dataset obtained from the Gene Expression Omnibus (GEO). The analysis was performed using the GEOquery package in R. This dataset comprised gene expression of EMT in PTC invasion, including 18 samples (four normal tissues and 14 thyroid cancer tissues). We employed the limma method to identify differentially expressed genes associated with EMT. We considered significance based on an adjusted p-value (Adjpval) threshold of less than 0.01 and an absolute log fold change (logFC) greater than 1.
Selection of LPAR1 as the main study target
We adopted a comprehensive approach by intersecting the results of RNA expression analysis obtained from the TCGA-THCA dataset with those derived from the GEO dataset. Through this analysis, we identified LPAR1 as a target of interest for further investigation and analysis.
Protein-protein interaction (PPI) analysis
To better understand LPAR1 protein-protein interactions and their potential to relate to EMT, we utilized the GeneMANIA database. GeneMANIA serves as a valuable resource for exploring functional associations between genes. Through this powerful database, we explored the protein-protein interactions between LPAR1 and its relevant partners. The GeneMANIA database integrates diverse sources of applicable association data, including genetic interactions, co-expression patterns, pathways, and physical interactions. We aimed to uncover a more comprehensive view of the interactions within the protein-protein interaction (PPI) network involving LPAR1 by harnessing GeneMANIA database capabilities.
Functional enrichment analysis of PPI network
To gain a deeper understanding of the functions and enriched pathways associated with the proteins identified in the protein-protein interaction (PPI) network, we performed gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses. The GO analysis encompasses 3 aspects: cellular component (CC), biological process (BP), and molecular function (MF). To conduct the GO analysis, we utilized the ClusterProfiler R package (Version 4.0.2) and employed the Benjamini-Hochberg method with a significance threshold of P-value < .05 and q-value < .05. This allowed us to identify the functional categories and molecular functions associated with the proteins. Additionally, the KEGG analysis revealed the potential roles of these genes within specific pathways.
Immune infiltration analysis
To investigate the potential correlation between LPAR1 expression and immune infiltrates in PTC, we utilized the Tumor Immune Estimation Resource (TIMER). This online tool leverages TCGA data to comprehensively analyze immune infiltrates across various cancer types. We computed the abundance of immune infiltrates, including CD8+ T cells, CD4+ T cells, dendritic cells, neutrophils, B cells, and macrophages, within PTC samples using partial deconvolution and linear least square regression. We aimed to establish a connection between LPAR1 expression levels and immune infiltrates in PTC samples. This was done by exploring potential associations between LPAR1 and the immune microenvironment in PTC.
Evaluation of LPAR1 expression in various cancers through the Gepia2 database
To expand our knowledge of LPAR1 expression across various types of cancer, we utilized the Gepia2 database (http://gepia2.cancer-pku.cn/). Gepia2 is a valuable tool for large-scale RNA sequencing and expression analysis. This database integrates data from TCGA and the Genotype-Tissue Expression (GTEx) projects. By accessing the Gepia2 database, we explored and examined LPAR1 expression levels in various cancer types.
Prediction of miRNAs related to LPAR1
Our study utilized the multiMiR tool in the R programing language to predict miRNA target genes and investigate miRNA-mRNA interactions. The multiMiR tool, available through its web server at http://multimir.org/, consolidates a wide range of predicted and validated miRNA-target interactions. It incorporates numerous databases, providing a comprehensive selection of interactions based on various binding strength thresholds, including those associated with diseases and medications. Our research focused on the TarBase database within multiMiR, which offers verified miRNA-target interactions.
Cell culture
The Nthy-ori 3-1 human thyroid epithelial cell line was obtained from the American Type Culture Collection (Manassas, VA, USA), while the BCPAP human papillary thyroid carcinoma cell line was obtained from the Chinese Academy of Sciences. Both cell lines were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen Technologies, USA) supplemented with 10% fetal bovine serum (FBS), 100 U/mL penicillin, and 100 µg/mL streptomycin. The cells were incubated at 37°C in a humidified atmosphere with 5% CO2.
RNA isolation, cDNA synthesis, and RT-qPCR
Cell lines were subjected to RNA isolation using TRIzol (Life Technologies, USA) following the manufacturer’s instructions. Subsequently, RNA was reverse transcribed into cDNA using the PrimeScript RT kit (Takara Biotechnology). The expression of LPAR1 was assessed by RT-qPCR on a Light Cycler 480II real-time PCR system (Roche Diagnostics, Switzerland) using SYBR Premix Ex Taq™ (Takara, Dalian, China) as the detection method, with GAPDH serving as the endogenous control. The 2−∆∆Cq method was utilized to determine the relative expression values, enabling analysis of the gene expression changes observed in the real-time quantitative PCR experiments. The primer sequences employed in our study are provided below: LPAR1 (F: 5’- CTCATTGACACCACCGTGAC-3’; R: 5’-GTTGAAAATGGCCCAGAAGA-3’), GAPDH (F: 5’-GTCTCCTCTGACTTCAACAGCG -3’; R: 5’-ACCACCCTGTTGCTGTAGCCAA-3’).
Statistical analysis
The data were analyzed using GraphPad Prism 5.0 statistical software (GraphPad Software, Inc., La Jolla, CA, USA). Statistical differences between groups were evaluated using an unpaired t-test. A significance level of P < .05 was used to determine statistical significance. To ensure robustness and reliability, all experiments were performed in duplicate with at least 3 replications, which minimized the chance of chance variations, thus strengthening the validity of the findings.
Results
Identification of differentially expressed mRNAs from GEO database pertaining to EMT
Using the limma package, we analyzed the GSE6004 dataset obtained from the GEO database. We identified differentially expressed genes (DEGs) that distinguish primary tumors from normal tissues. Through this analysis, we identified a total of 40 upregulated genes and 28 downregulated genes. The results are visually represented in the Volcano map (Figure 1a).
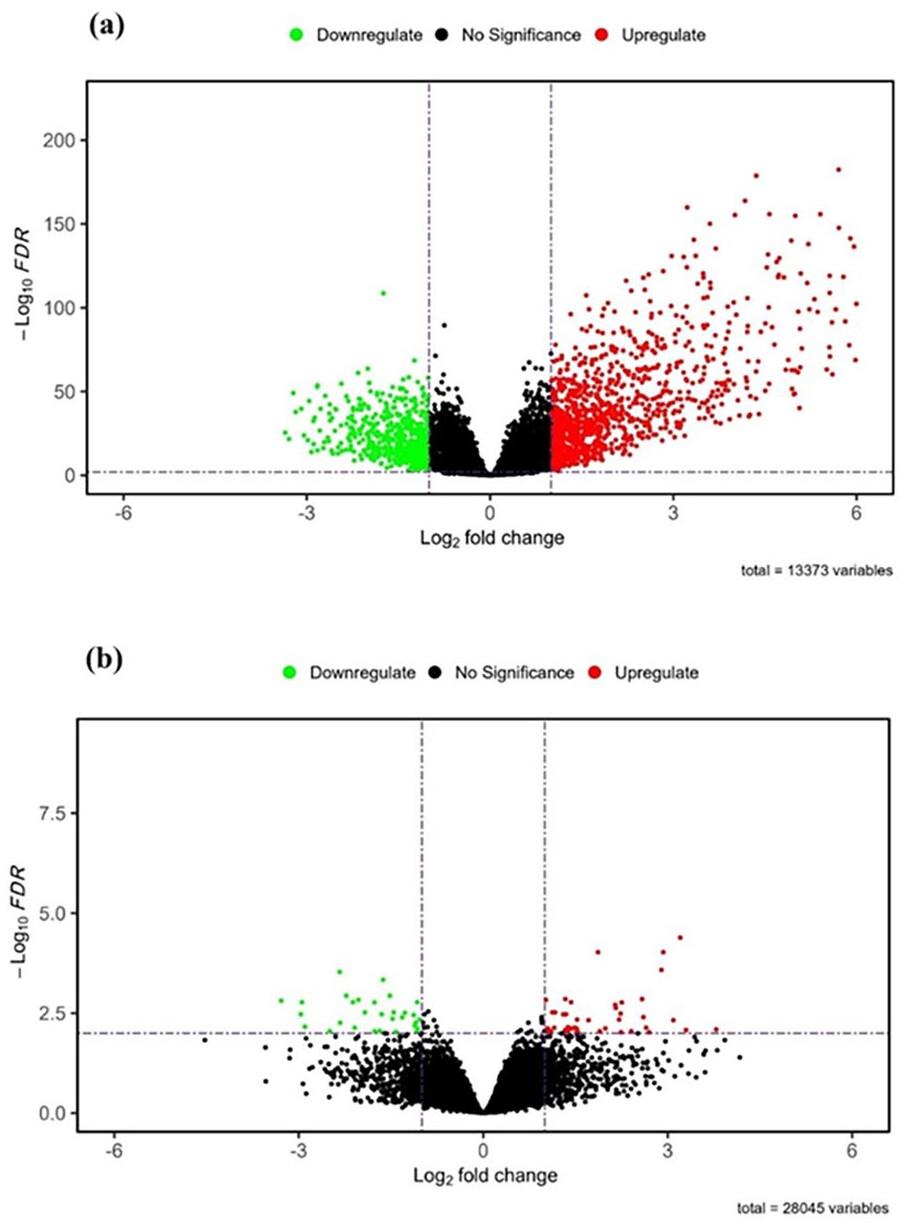
Identification of differentially expressed mRNAs from the GDC database
By employing the DESeq2 package in R, we conducted an analysis to identify a set of differentially expressed genes (DEGs) in thyroid cancer samples compared to normal control samples. Our analysis revealed a total of 1081 upregulated genes and 544 downregulated genes, indicating significant changes in gene expression between the 2 groups (Figure 1b).
Identification of genes with implications in EMT
To gain insights into the genes associated with EMT, we conducted a comprehensive analysis by comparing the DEGs obtained from the TCGA results with those derived from the GEO dataset. Through this meticulous process, we identified a set of 44 genes common to both datasets. These 44 genes underwent a comprehensive analysis to assess their significance. Among them, LPAR1 emerged as the most compelling candidate, displaying significant potential and warranting further in-depth investigation. LPAR1 is a promising gene for elucidating its role in EMT and potential implications for related processes (Table 1) (Figure 2) (Figure 3).
| TCGA LPAR1 expression results | GSE6004 LPAR1 expression results | ||||
|---|---|---|---|---|---|
| logFC | P-value | FDR | logFC | P-value | Adj. P-value |
| −1.32 | 6.58E-30 | 8.02E-29 | −1.43 | 2.83E-05 | .009548 |
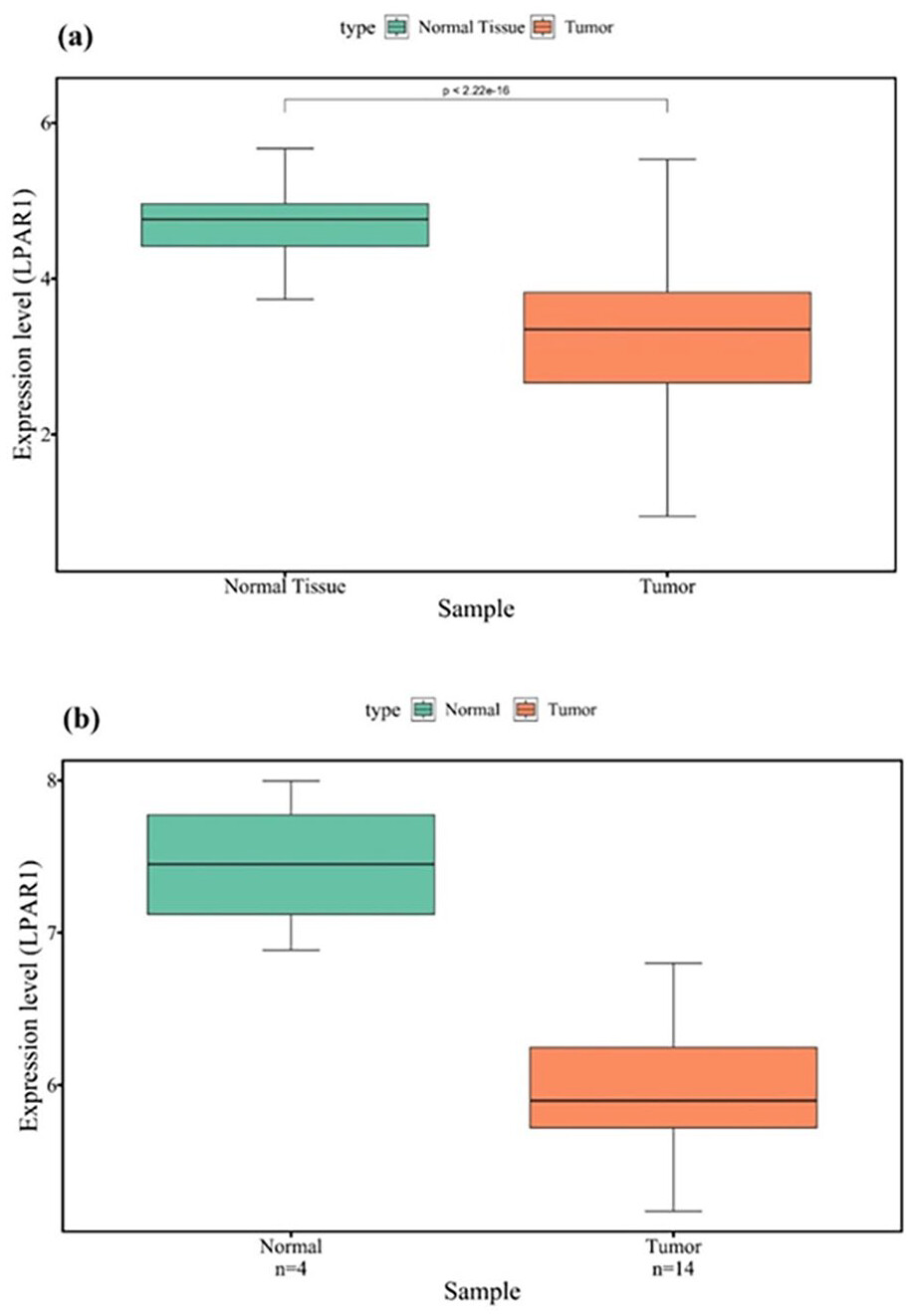
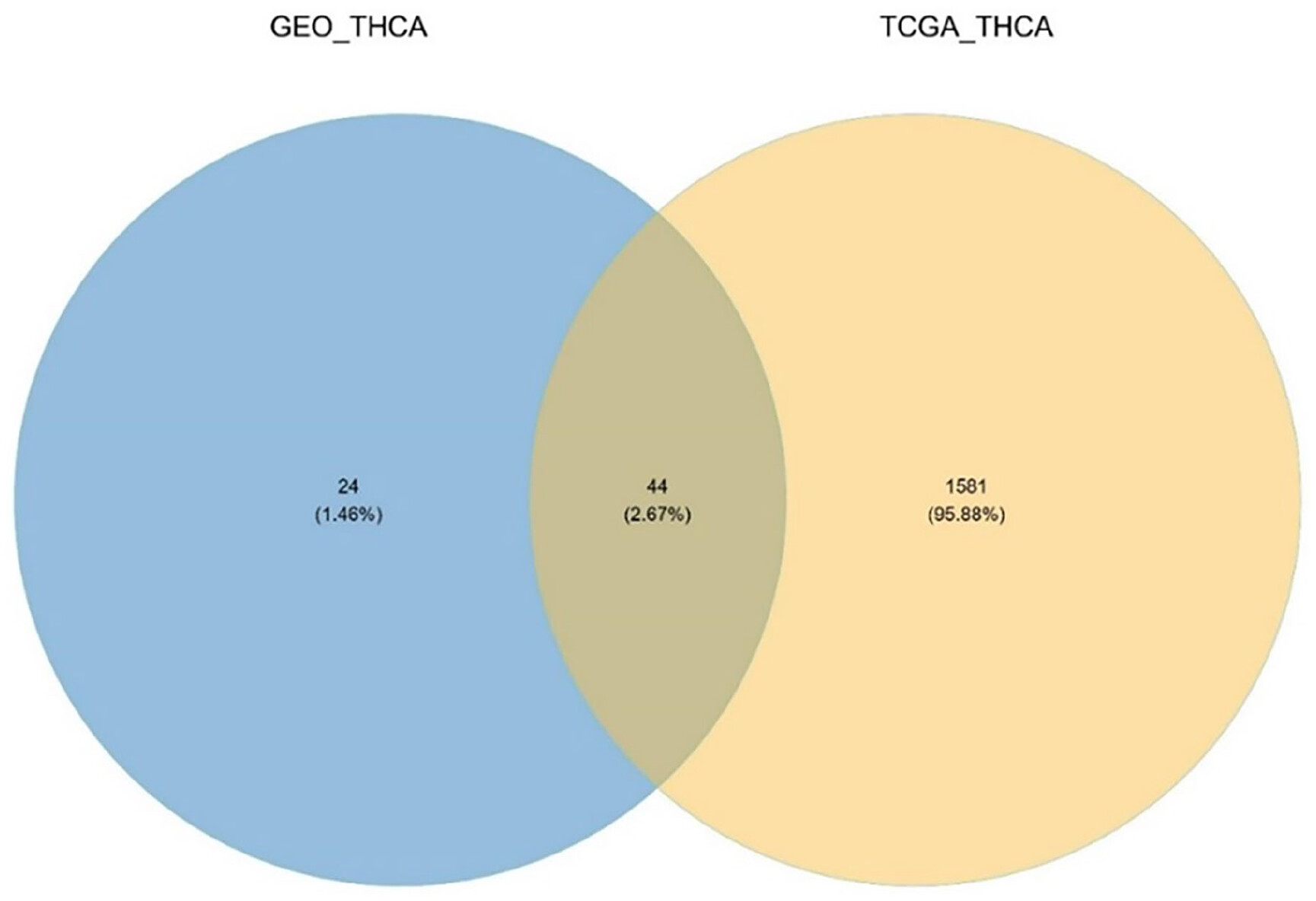
Protein-protein interaction (PPI) analysis of LPAR1
To explore the intricate protein-protein interactions involving LPAR1, we utilized the GeneMANIA plugin within Cytoscape. Through our analyses, we generated detailed networks that visually represented these interactions. To ensure the accuracy and reliability of our representation, we incorporated distinct data sources for each edge in the network.
Employing the GeneMANIA database integrated within Cytoscape, we delved into the top 20 proteins related to LPAR1 based on co-expression, co-localization, genetic interactions, pathways, physical interactions, predicted interactions, and shared protein domains. Our analysis revealed a network consisting of 21 nodes and 115 edges. Within this network, several pivotal proteins were closely associated with LPAR1. Notable mentions include GNA13, GNA12, LPAR2, ARHGEF12, PTK2, PTK2B, LPAR3, ARHGEF11, GAB1, AKAP13, ITGB4, S1PR3, CXCL8, PLD2, NFKBIA, CASP3, GNG2, ENPP2, MMP2, and PDE1B. These proteins are intricately linked to LPAR1, suggesting potential functional relationships and providing insights into the protein network associated with LPAR1 (Figure 4).
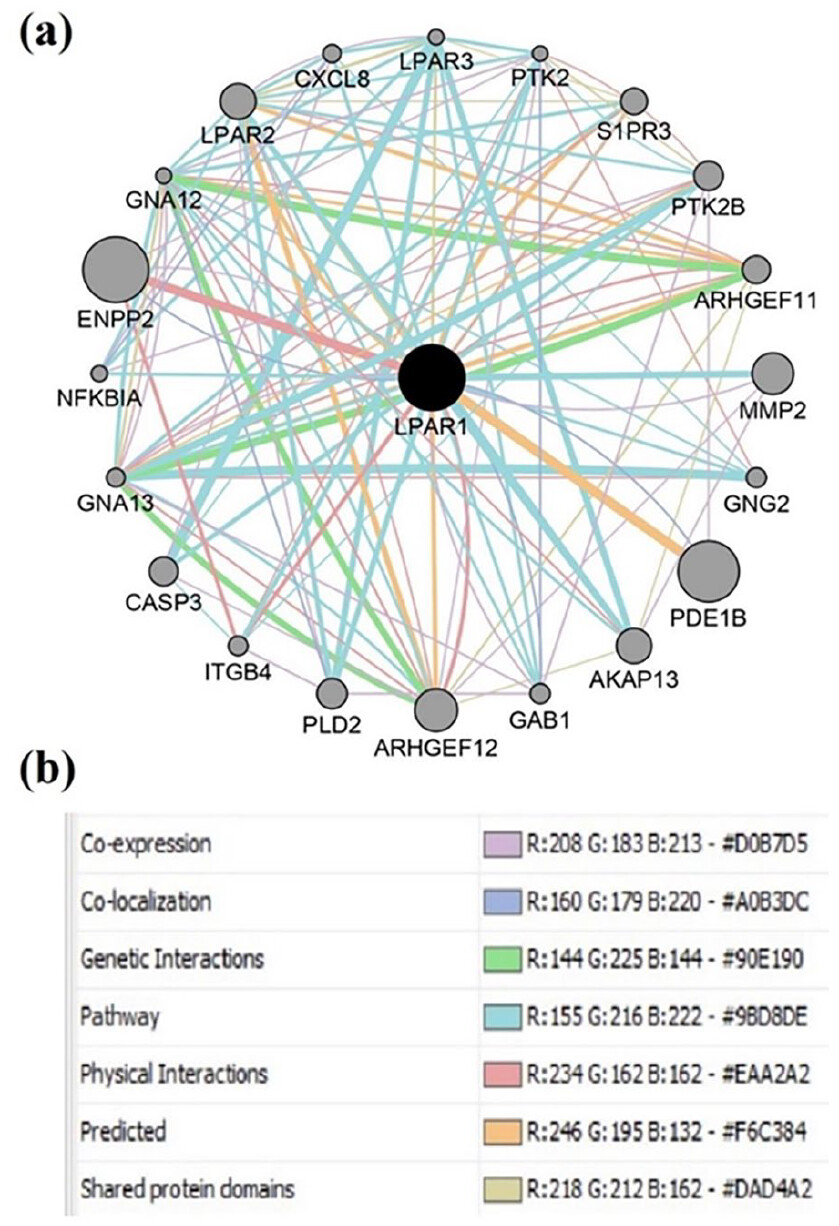
Functional enrichment analysis of DEGs
We conducted GO and KEGG enrichment analyses to understand the biological significance and signaling pathways associated with genes in the LPAR1 protein-protein interaction (PPI) network. Based on the analysis of biological processes (BP), the top 3 GO outcomes underscored associations with “Rho protein signal transduction,” “small GTPase-mediated signal transduction,” “positive regulation of cytosolic calcium ion concentration,” and “Ras protein signal transduction” (Figure 5). The cellular component (CC) aspect highlighted enrichment in the “heterotrimeric G-protein complex,” “GTPase complex,” and “focal adhesion” (Figure 6). The molecular function (MF) domain emphasized genes enriched in “bioactive lipid receptor activity,” “G protein-coupled receptor binding,” and “guanyl-nucleotide exchange factor activity”(Figure 7). Notably, KEGG pathway analysis revealed prominence in “Human cytomegalovirus infection,” “Phospholipase D signaling pathway,” “Pathogenic Escherichia coli infection,” “Regulation of actin cytoskeleton,” and “Parathyroid hormone synthesis, secretion, and action”(Figure 8).




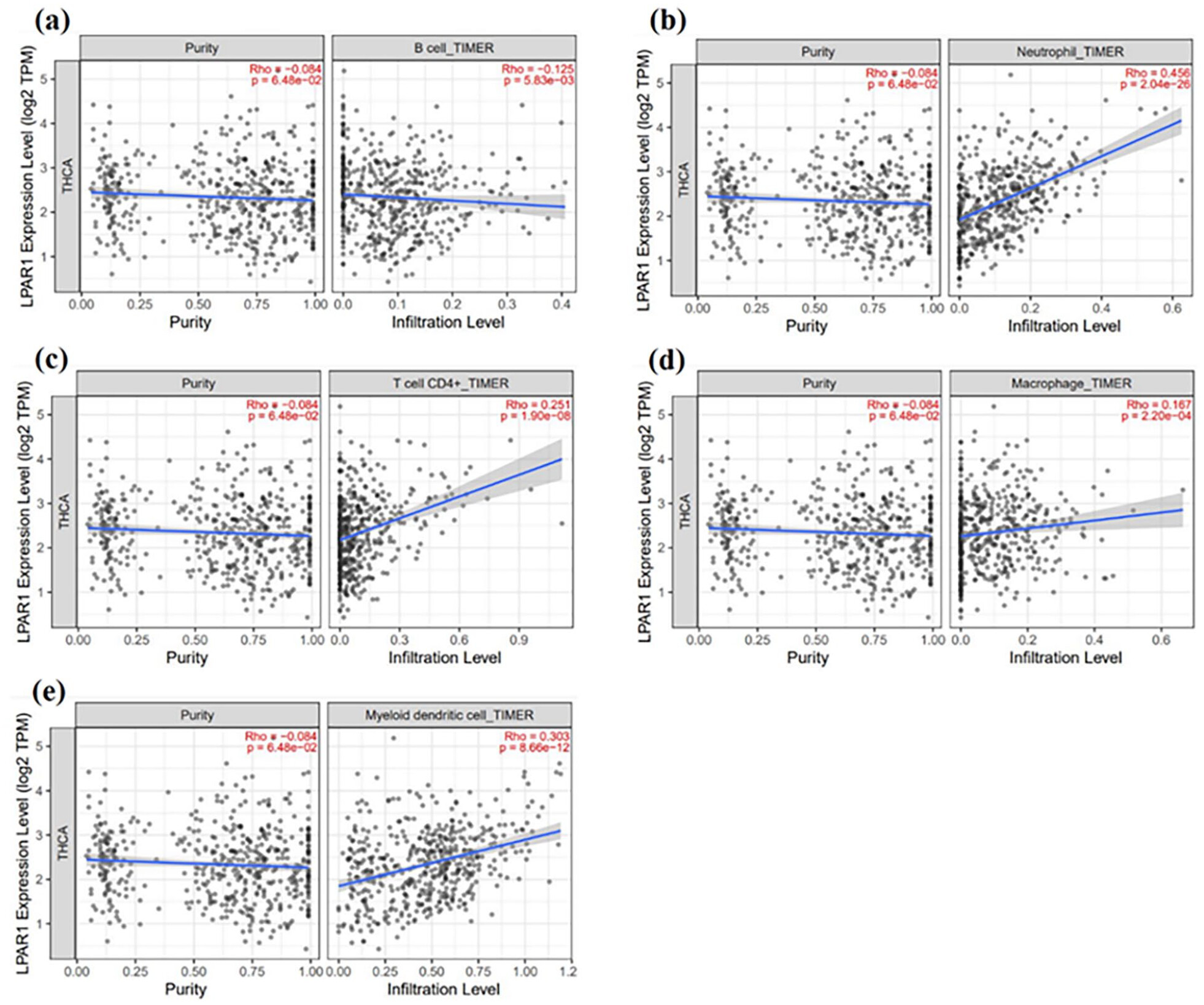
Correlation of LPAR1 expression with immune infiltration
We examined the relationship between LPAR1 expression and immune cell abundance to recognize the crucial role of immune infiltration in cancer. Within the tumor microenvironment, immune cells such as B and T lymphocytes, macrophages, neutrophils, and dendritic cells play pivotal roles. Using the TIMER tool, we conducted a meticulous analysis to scrutinize this correlation. The results exhibited positive correlations between LPAR1 expression and the infiltration of T cell CD4+ (Rho = 0.251, P = 1.90e−08), macrophages (Rho = 0.167, P = 2.20e−04), neutrophils (Rho = 0.456, P = 2.04e−26), and myeloid dendritic cells (Rho = 0.303, P = 8.66e−12). However, no significant correlation was found between LPAR1 expression and CD8+ T cell infiltration (Rho = −.024, P = 5.95e−01). Additionally, an inverse correlation was observed between LPAR1 expression and B cell infiltration (Rho = −.125, P = 5.83e−03; Figure 9).
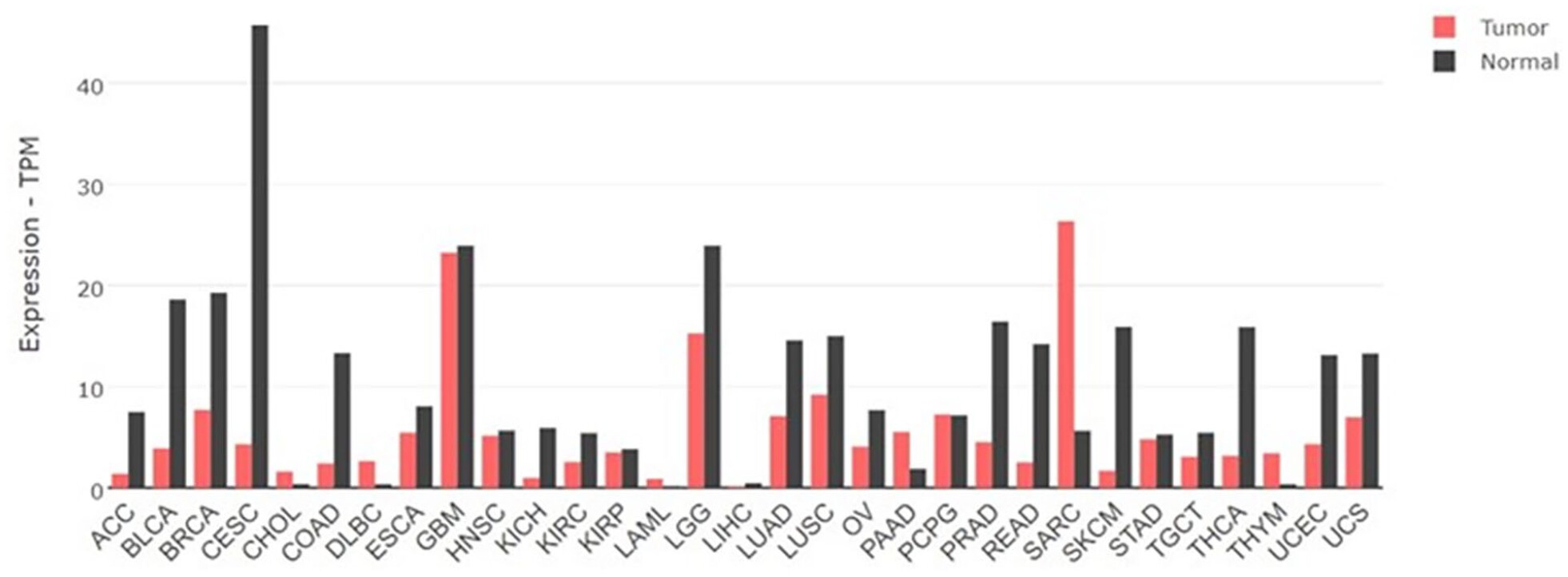
LPAR1 expression in various cancers
Our comprehensive analysis investigated LPAR1 expression patterns in 31 cancer types. Our findings revealed significant downregulation of LPAR1 expression compared to normal tissues in several cancer types, specifically in Adrenocortical carcinoma (ACC), Bladder Urothelial Carcinoma (BLCA), Cervical squamous cell carcinoma, endocervical adenocarcinoma (CESC), Colon adenocarcinoma (COAD), Kidney Chromophobe (KICH), Prostate adenocarcinoma (PRAD), Rectum adenocarcinoma (READ), Skin Cutaneous Melanoma (SKCM), Uterine Corpus Endometrial Carcinoma (UCEC), and Thyroid carcinoma (THCA). These observations provide valuable insights into the potential involvement of LPAR1 in the development and progression of these specific cancer types (Figure 10).
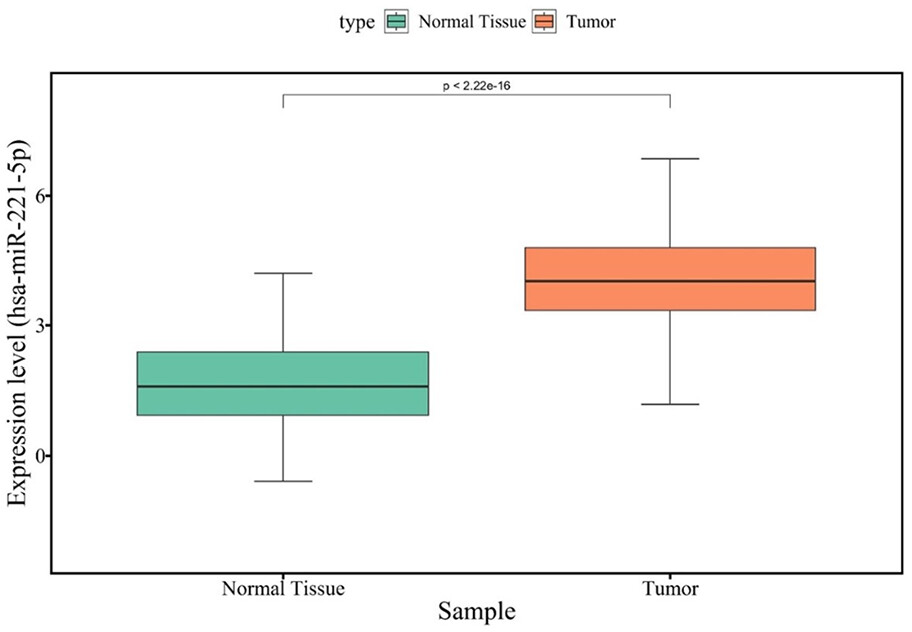
Prediction of miRNA-mRNA interactions
We utilized the powerful multiMiR tool in the R programing language to explore miRNA-mRNA interactions. This resource allowed us to uncover relationships between miRNAs and their target genes, including LPAR1. By analyzing miRNA binding sites on LPAR1, we predicted regulatory interactions within this intricate network. Among the identified interactions, we discovered a relationship between miR-221-5p and LPAR1. This interaction was further supported by experimental evidence found in the TarBase database, validating its significance. The miR-221-5p/LPAR1 interaction is evidence of the complex regulatory mechanisms that govern LPAR1 influence (Figure 11).
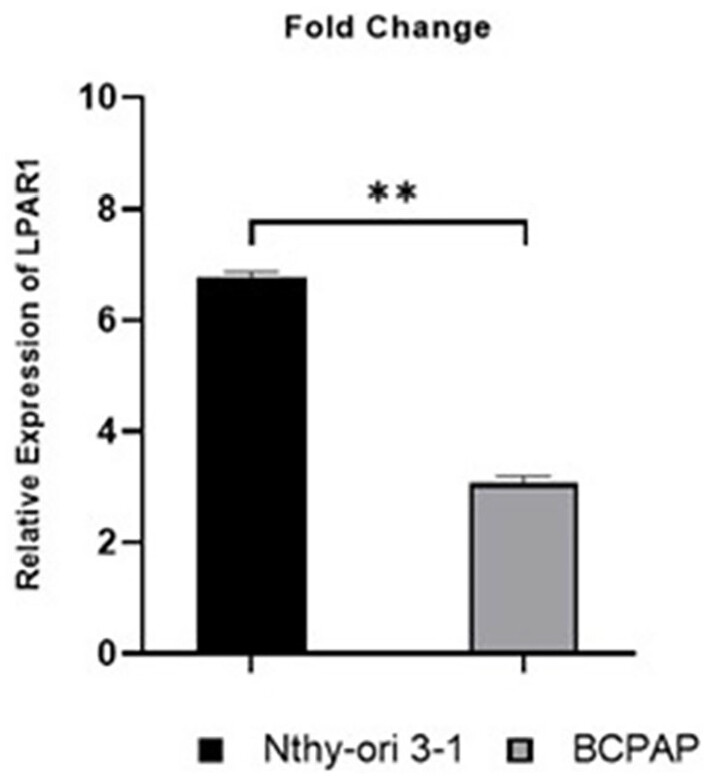
LPAR1 is downregulated in PTC cell line
To investigate the potential involvement of LPAR1 in PTC, we examined the relative expression of LPAR1 in 2 cell lines. The results revealed that the average expression of LPAR1 was significantly decreased in the PTC cell line BCPAP compared to the normal thyroid cell line Nthy-ori 3–1 (P < .01, Figure 12). The data also indicated a noticeable trend wherein more advanced and metastatic stage cancers exhibited lower levels of LPAR1 expression.
Discussion
The onset of EMT in thyroid epithelial cells results in the aberrant regulation of multiple factors, such as transcription factors, signaling cascades, epigenetic changes, and the tumor environment. These dysregulations significantly contribute to the recurrence and development of metastasis.22,23 Existing literature has demonstrated that disrupted LPAR1 expression plays a role in promoting tumor metastasis through diverse mechanisms. For instance, phosphatidic acid (PA) has been shown to enhance cell migration by interacting with LPAR1, leading to the activation of signaling molecules such as MAPKs (ERK1-2, p38, and JNK). Additionally, it triggers the upregulation of the JAK2/STAT3 pathway.24 In a study by Ha et al, it was revealed that LPA employs a signaling pathway mediated by Gαi2 and involves Src kinase to increase the levels of HIF1α and specific factors associated with EMT, like Slug. This process contributes to the invasive migration of ovarian cancer cells.25 Furthermore, LPA induces EMT in ovarian tumors by activating the Wnt/β-catenin signaling pathway, which relies on the presence of β1-integrin. This novel mechanism provides a means for activating this pathway independent of mutations, contributing to the progression of epithelial ovarian cancer.26 However, the involvement of LPAR1 in EMT and its contribution to promoting EMT in the context of PTC remain unknown. It is crucial to comprehend the role, expression pattern, and regulatory mechanisms of LPAR1 in PTC pathogenesis to establish a foundation for targeting this pathway therapeutically. This study aimed to investigate LPAR1 as a significant regulator of human thyroid cancer cell dissemination and propose that targeting LPAR1 could offer novel strategies for reducing malignancy in PTC.
Our study aimed to identify DEGs between PTC and control cells to understand PTC pathogenesis and uncover diagnostic biomarkers and therapeutic targets, focusing on EMT. Through analysis, we identified LPAR1 and confirmed its lower expression in PTC cells. Further research revealed the involvement of the has-miR-221-5p/LPAR1 network in PTC cell EMT. The research findings on LPAR1 highlight its diverse signaling pathways and functions in different tumor cell types. Notably, LPAR1 has been reported as a negative cell motility and invasion regulator, promoting epithelial maturation.27 In prostate cancer, LPAR1 via the downstream Gα pathways plays a role in facilitating multicellular differentiation while suppressing invasiveness. Clinical data analysis further supports that LPAR1 and Gα12/13 signaling act as metastasis suppressors, regulating the balance between cell motility, invasion, and epithelial maturation.12
LPAR1 exhibited substantial downregulation in gastric cancer tissues, displaying an inverse association with the expression of miR-501-5p. Notably, the reduction in LPAR1 expression was found to enhance both cell proliferation and migration. However, these effects were counteracted when gastric cancer cells were co-transfected with a miR-501-5p inhibitor. These findings suggest that miR-501-5p can promote cell proliferation and migration in gastric cancer by downregulating LPAR1. Therefore, the miR-501-5p/LPAR1 axis holds promise as a potential therapeutic target for the treatment of gastric cancer.18 Conversely, our findings contrast with previous studies that indicate an elevation in LPAR1 expression in certain tumors, leading to tumor-promoting effects. For instance, it has been demonstrated that heightened levels of LPAR1 stimulate increased HIF1α levels through the Gαi2-mediated signaling pathway involving Src kinase. This, in turn, triggers EMT and facilitates invasive cell migration, accompanied by upregulated expression of N-cadherin, Slug/Snail2 in ovarian cancer cells.25 Additionally, LPAR1 inhibition effectively prevented pulmonary metastasis of osteosarcoma cells.28
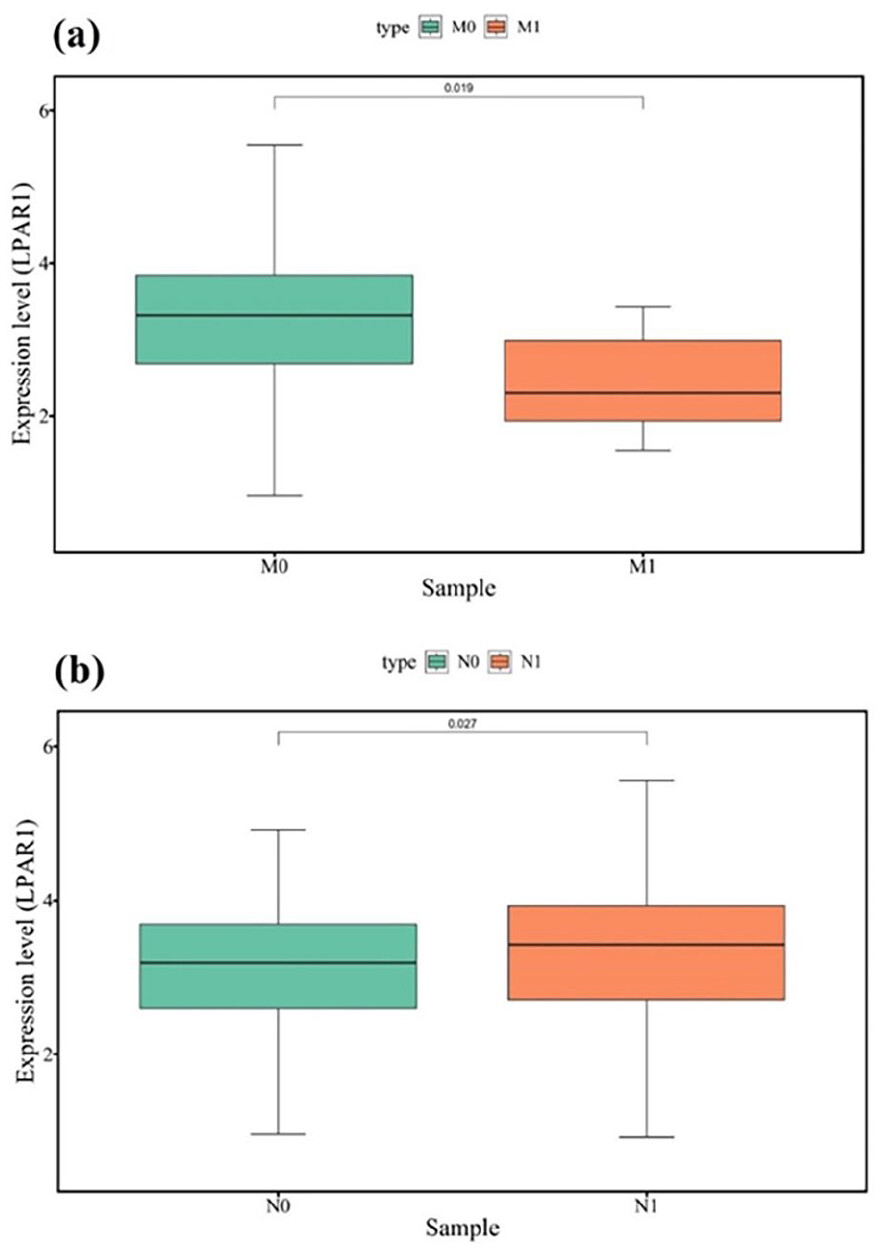
Our bioinformatics-based study underscores the importance of LPAR1 in the EMT of thyroid cancer. We observed downregulation of LPAR1 expression in PTC tissue compared to normal tissue. Our analyses have revealed a decrease in LPAR1 expression among PTC patients with distant metastasis compared to those without metastasis. Interestingly, despite the overall reduced expression levels of LPAR1 in PTC, we observed higher expression in patients with lymph node metastasis than those without lymph node involvement. This suggests that reduced LPAR1 levels can serve as a distinguishing factor, differentiating between PTC patients with lymph node metastasis and distant metastasis and distinguishing metastatic from non-metastatic patients (Figure 13).
RT-qPCR experiment corroborated the findings of our in-silico analyses, demonstrating a significant downregulation of LPAR1 expression in the PTC cell line compared to the normal cell line. Additionally, significant differences in LPAR1 expression were observed between different stages of PTC, including stage I versus stage II and stage II versus stage III. Our investigation also revealed the involvement of LPAR1 in metastasis-related pathways, including phospholipase D signaling and actin cytoskeleton regulation. We found compelling evidence of LPAR1’s interaction with key proteins associated with EMT in PTC, such as ITGB4, GAB1, GNA13, ARHGEF12, PTK2, and CXCL8. The tumor microenvironment (TME) plays a pivotal role in cancer pathogenesis, which exerts its influence by modulating the balance between suppressive and cytotoxic responses in the vicinity of the tumor, thereby impacting disease outcomes.29,30 However, the specific role of LPAR1 within the TME remains poorly understood. Using the TIMER database, we investigated the impact of LPAR1 on tumor-infiltrating immune cells in PTC. Notably, LPAR1 shows positive associations with antitumor immune cells, including effector CD4+ T cells, macrophages, neutrophils, and dendritic cells. Conversely, there is a negative correlation between LPAR1 expression and tumor-infiltrating B lymphocytes, which is consistent with a study by Shi et al, who explored the role of LPAR1 in prostate cancer.31,32
The interaction between LPAR1 and miRNAs plays significant roles in various cancers’ pathophysiology.33 For instance, miR-892b negatively regulates LPAR1 expression, and downregulation of miR-892b with simultaneous upregulation of LPAR1 enhances proliferation, migration, and invasion of nasopharyngeal carcinoma cells.34,35 Our analysis predicts LPAR1 as a potential target of miR-221-5p, which is significantly overexpressed in PTC and inversely correlated with LPAR1 expression. The upregulation of miR-221-5p potentially suppresses LPAR1, promoting EMT in PTC. Previously, it was reported that miR-221/222 clusters are significantly upregulated in PTC compared to adjacent normal tissue. Increased levels of miR-221/222 are linked to tumor size, TNM stage, lymph node metastasis, and extra-thyroidal extension. Moreover, miR-221/222 promotes migration and invasion while inhibiting apoptosis and autophagy in PTC cells by targeting ATG10.36 In contrast, miR-221-5p has been found to play a tumor-suppressive role in patients with prostate cancer. Loss-of-function and gain-of-function experiments have provided evidence that the overexpression of miR-221-5p reduces cell proliferation, migration, and EMT markers in vitro. Moreover, it leads to decreased tumor growth, cell extravasation, and formation of metastases in vivo.37
In summary, our study utilizing bioinformatic analyses demonstrates that LPAR1 acts as a metastasis-suppressor in PTC. Decreased expression of LPAR1 is associated with promoting EMT and metastatic behavior in PTC patients. Additionally, LPAR1 expression correlates with tumor-infiltrating immune cells, suggesting its potential as an immune-related biomarker in PTC. This highlights the potential role of LPAR1 as a prognostic biomarker for EMT and metastasis in PTC. Moreover, the findings from our study propose LPAR1 as a potential therapeutic target in PTC. As metastasis-suppressor, restoring LPAR1 expression or manipulation of its downstream signaling pathways may have therapeutic benefits in inhibiting the development of metastatic phenotype and improving patient outcomes. Hence, identifying patients with altered LPAR1 expression levels could aid in predicting their response to immunotherapeutic interventions and guiding treatment decisions. To gain a deeper understanding of the mechanisms through which LPAR1 contributes to the metastasis of PTC, further experiments conducted both in vitro and in vivo are required.
Conclusion
Our bioinformatics-based study demonstrates that LPAR1 functions as a metastasis-suppressing factor and is closely associated with EMT. The observed decrease in the expression of LPAR1, coupled with the overexpression of miR-221-5p in tumor cells, provides supporting evidence for the role of LPAR1 in suppressing metastasis. Additionally, the analyses highlight the potential of LPAR1 as a valuable biomarker for diagnosis and prognosis. These findings have important implications for understanding PTC pathogenesis and contributes to the development of novel therapeutic strategies for patients with PTC.
Acknowledgments
Not applicable.
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
ORCID iD
Mohammad Natami https://orcid.org/0009-0009-6915-487X
Data availability statement
The datasets used in the current study are available from the corresponding author on reasonable request.
References
1. Xie Z, Lun Y, Li X, et al. Bioinformatics analysis of the clinical value and potential mechanisms of AHNAK2 in papillary thyroid carcinoma. Aging. 2020;12:18163-18180.
2. Zamora-Ros R, Cayssials V, Clèries R, et al. Sweetened beverages are associated with a higher risk of differentiated thyroid cancer in the EPIC cohort: a dietary pattern approach. Eur J Nutr. 2023;62:105-114.
3. Zamora-Ros R, Cayssials V, Franceschi S, et al. Polyphenol intake and differentiated thyroid cancer risk in the European Prospective Investigation into Cancer and Nutrition (EPIC) cohort. Int J Cancer. 2020;146:1841-1850.
4. Silva SN. Special Issue: Genetic Perspectives in Thyroid Cancer. Genes. 2021;12(2):126.
5. Kamali MJ, Salehi M, Mostafavi M, et al. Hijacking and rewiring of host CircRNA/miRNA/mRNA competitive endogenous RNA (ceRNA) regulatory networks by oncoviruses during development of viral cancers. Rev Med Virol. 2024;34(2):e2530.
6. Kyrodimos E, Chrysovergis A, Mastronikolis N, et al. The landscape of single nucleotide polymorphisms in papillary thyroid carcinoma. Cancer Diagn Progn. 2023;3:26-30.
7. Ywata de, Carvalho A, Kohler HF, Gomes CC, Vartanian JG, Kowalski LP. Predictive factors for recurrence of papillary thyroid carcinoma: analysis of 4,085 patients. Acta Otorhinolaryngol Ital. 2021;41:236-242.
8. Khare SK, Bajaj V. Optimized tunable Q wavelet transform based drowsiness detection from electroencephalogram signals. IRBM. 2022;43:13-21.
9. Lin JS, Bowles EJA, Williams SB, Morrison CC. Screening for thyroid cancer: updated evidence report and systematic review for the US Preventive Services Task Force. JAMA. 2017;317:1888-1903.
10. Abdullah MI, Junit SM, Ng KL, et al. Papillary thyroid cancer: genetic alterations and molecular biomarker investigations. Int J Med Sci. 2019;16:450-460.
11. Ghorbani M, Sadrian SH, Ghaderpanah R, et al. Tranexamic acid in total hip arthroplasty: an umbrella review on efficacy and safety. J Orthop. 2024;54:90-102.
12. Härmä V, Knuuttila M, Virtanen J, et al. Lysophosphatidic acid and sphingosine-1-phosphate promote morphogenesis and block invasion of prostate cancer cells in three-dimensional organotypic models. Oncogene. 2012;31:2075-2089.
13. Liu X, Pei M, Yu Y, Wang X, Gui J. Reduction of LPAR1 expression in neuroblastoma promotes tumor cell migration. Cancers. 2022;9;14(14):3346.
14. Sumitomo A, Siriwach R, Thumkeo D, et al. LPA induces keratinocyte differentiation and promotes skin barrier function through the LPAR1/LPAR5-RHO-ROCK-SRF axis. J Investig Dermatol. 2019;139:1010-1022.
15. Guerrero JA, Rivera J, Quiroga T, et al. Novel loci involved in platelet function and platelet count identified by a genome-wide study performed in children. Haematologica. 2011;96:1335-1343.
16. Rahimian Z, Feili A, Ghaderpanah R, et al. COVID-19-associated acute pancreatitis: a systematic review of case reports. Przegl Epidemiol. 2023;77:66-73.
17. Lin YH, Lin YC, Chen CC. Lysophosphatidic acid receptor antagonists and cancer: the current trends, clinical implications, and trials. Cells. 2021;10(7):1629.
18. Ma X, Feng J, Lu M, et al. microRNA-501-5p promotes cell proliferation and migration in gastric cancer by downregulating LPAR1. J Cell Biochem. 2020;121:1911-1922.
19. Kato K, Yoshikawa K, Tanabe E, et al. Opposite roles of LPA1 and LPA3 on cell motile and invasive activities of pancreatic cancer cells. Tumour Biol. 2012;33:1739-1744.
20. Memarian A, Saberi SM, Aghakhani K, et al. Physicians’ Awareness Towards Forensic Psychiatry, a Survey of Physicians Who Work in an Iranian General Hospital. 2018;43(1):98-104.
21. Ribatti D, Tamma R, Annese T. Epithelial-mesenchymal transition in cancer: a historical overview. Transl Oncol. 2020;13:100773.
22. Shakib H, Rajabi S, Dehghan MH, et al. Epithelial-to-mesenchymal transition in thyroid cancer: a comprehensive review. Endocrine. 2019;66:435-455.
23. Salehi M, Kamali MJ, Rajabzadeh A, et al. tRNA-derived fragments: key determinants of cancer metastasis with emerging therapeutic and diagnostic potentials. Arch Biochem Biophys. 2024;753:109930.
24. Gomez-Larrauri A, Gangoiti P, Camacho L, et al. Phosphatidic acid stimulates lung cancer cell migration through interaction with the LPA1 receptor and subsequent activation of MAP kinases and STAT3. Biomedicines. 2023;11:1804.
25. Ha JH, Ward JD, Radhakrishnan R, et al. Lysophosphatidic acid stimulates epithelial to mesenchymal transition marker slug/snail2 in ovarian cancer cells via gαi2, src, and HIF1α signaling nexus. Oncotarget. 2016;7:37664-37679.
26. Burkhalter RJ, Westfall SD, Liu Y, Stack MS. Lysophosphatidic acid initiates epithelial to mesenchymal transition and induces β-catenin-mediated transcription in epithelial ovarian carcinoma. J Biol Chem. 2015;290:22143-22154.
27. Shida D, Watanabe T, Aoki J, et al. Aberrant expression of lysophosphatidic acid (LPA) receptors in human colorectal cancer. Laboratory Investigation. 2004;84:1352-1362.
28. Takagi S, Sasaki Y, Koike S, et al. Platelet-derived lysophosphatidic acid mediated LPAR1 activation as a therapeutic target for osteosarcoma metastasis. Oncogene. 2021;40:5548-5558.
29. Lv B, Wang Y, Ma D, et al. Immunotherapy: Reshape the tumor immune microenvironment. Front Immunol. 2022;13:844142.
30. Chew V, Toh HC, Abastado JP. Immune microenvironment in tumor progression: characteristics and challenges for therapy. J Oncol. 2012;2012:608406.
31. Shi J, Jiang D, Yang S, et al. LPAR1, correlated with immune infiltrates, is a potential prognostic biomarker in prostate cancer. Front Oncol. 2020;10:846.
32. Memarian A, Mostofizadeh R, Aghakhani K, et al. Investigating emergency nurses’ awareness on the differentiation between acute methanol and ethanol intoxication. Int J -med Toxicol Forensic -med. 2021;11:33532.
33. Rafiyan M, Abadi MHJN, Zadeh SST, et al. Lysophosphatidic acid signaling and microRNAs: new roles in various cancers. Front Oncol. 2022;12:917471.
34. Peng J, Liu F, Zheng H, Wu Q, Liu S. Long noncoding RNA ZFAS1 promotes tumorigenesis and metastasis in nasopharyngeal carcinoma by sponging miR-892b to up-regulate LPAR1 expression. J Cell Mol Med. 2020;24:1437-1450.
35. Farrokhi M, Taheri F, Farrokhi M, et al. Advancements and innovations in cancer management: a comprehensive perspective. Kindle. 2024;4:1-161.
36. Shen H, Lin Z, Shi H, et al. MiR-221/222 promote migration and invasion, and inhibit autophagy and apoptosis by modulating ATG10 in aggressive papillary thyroid carcinoma. 3 Biotech. 2020;10:339.
37. Kiener M, Chen L, Krebs M, et al. miR-221-5p regulates proliferation and migration in human prostate cancer cells and reduces tumor growth in vivo. BMC Cancer. 2019;19:627.
Cite
Cite
Cite
OR
Download to reference manager
If you have citation software installed, you can download citation data to the citation manager of your choice
Information, rights and permissions
Information
Published In
Article first published online: September 8, 2024
Issue published: January-December 2024
Keywords
Data availability statement
Data is available for this article. View more information
PubMed: 39253536
Authors
Author Contributions
All authors contributed to the study conception and design. Original draft preparation was conducted by BZ, ZD, and NM. Writing, reviewing and editing of the draft manuscript was completed by RF, MR, AM, and BP. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Metrics and citations
Metrics
Publication usage*
Total views and downloads: 1038
*Publication usage tracking started in December 2016
Publications citing this one
Receive email alerts when this publication is cited
Web of Science: 4 view articles Opens in new tab
Crossref: 8
- Prognostic value of LPAR1 expression and methylation in low-grade gliomas: a meta-analysis of TCGA and CGGA datasets and functional validation
- Relevance of AKT and RAS Signaling Pathways for Antibody–Drug Conjugate Immunotherapies in Acute Lymphoblastic Leukemia
- Enhanced Anticancer Efficacy of Selenium Nanoparticles Encapsulated in Niosomes: A Novel Therapeutic Strategy
- Static magnetic field promotes the doxorubicin toxicity effects on osteosarcoma cells
- Neurotrophic Factors in Multiple Sclerosis: a Review of the Mechanism
- Drug Delivery Options for Riluzole in the Treatment of Amyotrophic Lateral Sclerosis
- Role of Antioxidants in the Management and Treatment of Autism Spectrum: A Review of Pathways
- Theoretical and Experimental Study of an Electrokinetic Micromanipulator for Biological Applications
Figures and tables
Figures & Media
Tables
View Options
View options
PDF/EPUB
View PDF/EPUBAccess options
If you have access to journal content via a personal subscription, university, library, employer or society, select from the options below:
loading institutional access options
Alternatively, view purchase options below:
Access journal content via a DeepDyve subscription or find out more about this option.



