Introduction
Our dietary choices, lifestyles, and environment broadly impact the epigenome, its actions and build the foundations for optimal health and longevity. The study of environmental and nutritional impacts upon the epigenome, throughout replication, development, evolution and life stages can be defined as nutritional epigenetics.
1It has been well documented that dietary and lifestyle choices within populations influence human gene expression and the consumption of vitamins and minerals through supplementation and fortification have demonstrated modulation of gene expression through the interaction at the level of the epigenome.
2Over the recent decade, B vitamins have been a strong nutritional epigenetic research focus for the manipulation of methylation and the prevention of neural tube defects.
3 Today; gene-editing techniques lead the research focus showing great promise for chronic disease management.
4 However, this technique brings forth several ethical issues and possesses unknown evolutionary consequences.
5 Therefore, natural modulation of the epigenome may be a preferable, non-invasive, sustainable, personalized alternative for chronic disease management.
This research provides a fresh perspective on the function of methylation, explores the potential for the less understood epigenetic modifications, and evaluates nutritional epigenetics for the management of chronic diseases.
Epigenetics
The perception of epigenetics and its meaning has changed significantly over the past half-century.
6 In 1942, Embryologist Conrad Waddington proposed the name epigenotype to describe a complex of developmental processes that occur between genotype and phenotype.
7 In the early 1960s, Doskočil & Šorm identified the distribution of 5-methylcytosines of deoxyribonucleic acids (DNA)
8 and researchers of the Rockefeller Institute began studying the structure, function and modulatory effects of histones on the regulation of ribonucleic acid (RNA) synthesis.
9 Thirteen years later, Holliday and Pugh described DNA modification by methylation in development,
10 and following this, epigenetic research exploded.
In October 1990, the Human Genome Project commenced and was completed by April 2003. The $2.7 billion project gave researchers the ability to begin piecing together nature’s genetic blueprint.
11The modern understanding of epigenetic mechanisms is the regulation of gene expression, which results from modifications in chromatin structure, histone tails, and nucleotides without an alteration in the DNA sequence; meaning an inheritable change in phenotype without a change in genotype.
6 Until recently, the exploration of epigenetic mechanisms had its limitations due to the inaccessibility and affordability of genomic sequencing technologies.
11 Recent advances in technology have reduced the cost of human genome sequencing by 99.99%,
11 making genomic research accessible and allowing the exponential expansion of research in the industry, including nutritional epigenetics.
Genetic differences and epigenetic inflexibility resulting in abnormal gene expression, differential or aberrant methylation patterns account for the vast majority of diseases including cancer,
12 autoimmune disease,
13 obesity,
14 metabolic diseases,
14 and complex multi-system conditions such as fibromyalgia
15 and chronic fatigue syndrome.
16The Epigenome
Shortly after discovering the cell specificity of histones in 1950;
17 Allfrey and colleagues discovered structural modifications of acetylation or methylation to histones, regulated RNA synthesis.
9,18 Within the genome, a histone octamer is at the core of the chromatin’s nucleosome and the nucleosome is complete when coiled in DNA.
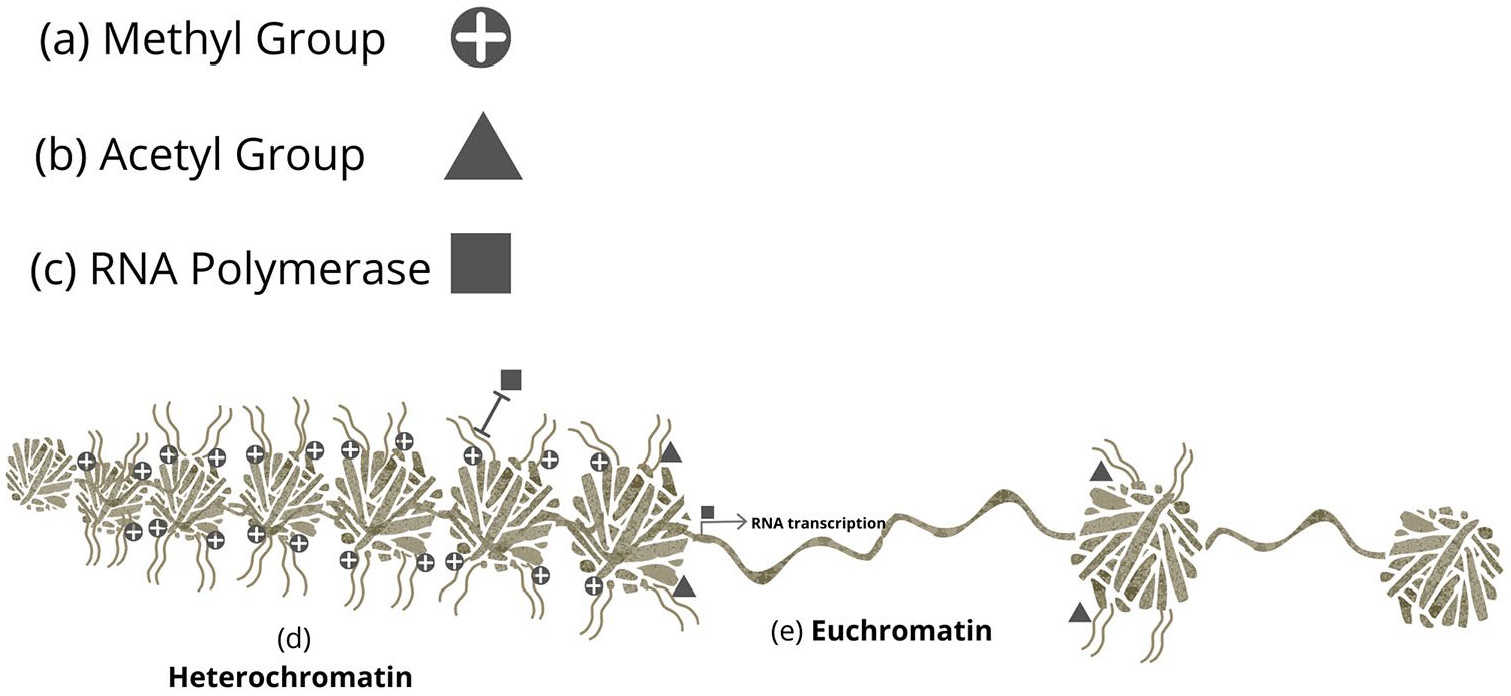
19 The histone tails that protrude from the nucleosome are the primary targets for structural changes to the epigenome.
19 The addition of a methyl or acetyl group at a specific amino acid residue upon the histone results in a conformational modification to the structure of the histone chromatin complex.
19 More recently, it has been demonstrated that histone tail residues can accept a variety of additional marks, including; phosphorylation, ubiquitination, sumoylation, citrullination, and glycosylation.
19 Histone tail modifications are identified by the histone number, the amino acid residue and the type of modification. For example; Histone 3/ Lysine 4/ dimethylation (H3K4me2).
19Nutritional Epigenetics
Human biochemistry is constantly evolving, with pathways frequently adapting to a changing environment.
20 Overtime; adaptations to changed environments result in subtle shifts in allele frequencies providing the foundations for evolution.
21,22Changes to epigenetic mechanisms as a result of environmental and dietary choices contribute to human physiology and biochemistry by altering gene expression.
23In medicine, manipulation of the epigenome is attractive as it can be rapid, reversible, specific, and capable of modulation beyond the blood-brain barrier (BBB).
24Vitamin, mineral, and phytochemical constituents derived from culinary foods have shown experimentally to have epigenomic modulatory capabilities with profound disease treatment potential.
25 The enzymatic activity for a variety of primary epigenetic histone modifiers are dependent on nutritionally derived cofactors such as nicotinamide adenonucleotide (NAD),
26 Zn2+,
26 ascorbate,
27,28 Fe2+,
29 endogenous tricarboxylic acid metabolites
30 and oxygen,
31 enabling specificity to nutritional modulation of the epigenome. Culinary, botanical herb and spice phytochemical constituents have demonstrated modulation of the epigenome in the laboratory also demonstrating disease treatment potential. However, transportation of a substance to a specific tissue is challenging.
32 In contrast; environmental contaminants have also demonstrated interaction at the epigenomic level influencing evolution and contributing to disease.
33 Collectively, research pertaining to alterations in gene expression and the resulting alterations to human biochemistry following short or long term exposure to exogenous dietary or environmental substance and the influence upon the structure and function of the epigenome can be defined as nutritional and environmental epigenetics.
1Rapid advances in technology have enabled a greater understanding of continual biological evolution and the bio-uniqueness of the human genome. Consequently, this requires unlearning and relearning of metabolic processes to keep up with the evolving biology, for the majority of metabolic pathways including research pertaining to nutrigenomics was originally undertaken without thorough consideration of the workings of the epigenome.
Old ideas and one carbon metabolism
Methylation has been a focus for nutritional epigenetic researchers since the mid-nineties when a group of leading researchers discovered associations between folate deficiency, 5,10 methylenetetrahydrofolate deficiency, methylenetetrahydrofolate reductase (MTHFR) inefficiency, neural tube defects, and cardiovascular disease.
34-41 Today, a PubMed search of the MTHFR enzyme at the National Library of Medicine generates over 7000 entries and has been associated with hundreds of conditions.
42 Single nucleotide variations (SNV) of MTHFR, the primary regulator of one-carbon metabolism and methyl group synthesis, was said to be the leading cause of active folate deficiencies leading to neural tube defects.
35B vitamins, including folate (B9) molecules derived from the diet, are considered cofactors for one-carbon metabolism, function as carriers of one-carbon methyl units
2 and are well known for the epigenetic modulation of methylation.
43 Analysis of metabolites and cofactors of one-carbon metabolism in erythrocytes, serum, and plasma, provide insight into vitamin deficiency, one-carbon metabolism activity and methylation.
2Fortification of grains, including wheat flour and corn maze for the prevention of folate deficiency, had been discussed since 1974,
44 but was not implemented in the United States until 1997 following repeated associations with folic acid depletion and the incidence of neural tube defects.
44DNA methylation
Methylation is a primary epigenetic mark essential for the regulation of gene expression and is generally associated with gene repression.
45 Methyl groups for endogenous methylation of histones, nucleotides, and proteins are said to be derived from one-carbon metabolism.
2 DNA methylation is a widely studied but not very understood epigenetic modification. DNA methyltransferase enzymes (DNMT) initiate DNA methylation through the binding of a methyl group to position 5′ of cytosine bases neighboring guanosine (CpG), generating 5-methyl-cytosine (5-mC). 5-mC is then oxidized to 5-hydroxymethylcytosine (5-hmC), which is said to promote de-methylation.
45 The Ten-Eleven Translocation di-oxygenase (TET) family of enzymes are shown to contribute to the removal of the methyl group from cytosine, forming 5-formylcytosine (5-fC) and 5-carboxylcytosine (5-caC).
45CpG’s are found in clusters which are referred to as islands, spanning approximately 1000 bp at the promoter region of a gene and are generally found unmethylated.
46 Methylated CpG’s at the promoter region of a gene are associated with gene repression.
45 Differential methylation regions (DMR) are regions upstream of gene promoters that also influence a gene’s expression.
475′ methylcytosines contribute to the rate of evolution as they are shown to deaminate at a much greater rate than unmethylated cytosines and are considered mutational hot spots within the genome.
48,49 Thymine DNA glycosylate (TDG) is essential for base excision repair throughout the DNA methylation and remethylating process.
45Promoter DNA methylation of any gene which results in a loss of gene function can have profound effects on phenotype and has the potential to cause disease.
50 Inhibition and gene knock out studies which result in loss of function can often predict phenotypical outcomes of DMR or promoter region hypermethylation.
47 For example; both promoter region CpG hypermethylation and DMR of the estrogen receptor1 (ESR1) which encodes the estrogen receptor alpha protein results in reduced ESR1 expression.
47,51 DMR of ESR1 has been implicated in a range of hormone-related diseases including ovarian endometriosis and resistance to hormone therapy.
47,51Tryptophan hydroxylase 2 (TPH2) is essential for the first rate-limiting step in the synthesis of the 5-hydroxytryptophan (5-HTP), the immediate precursor to serotonin.
52 In animal models, reduced TPH2 activity is associated with depression-related behavior, aggression and altered sexual preference.
53,54DNA methylation and reduced expression of TPH2 in humans has also been associated with suicide attempts in major depression.
52Histone methylation
The genome is comprised of hundreds of genes encoding writer histone methyltransferase enzymes capable of writing methylation to the histone.
19 Methylation of a repressive histone residue such as histone-3 lysine-9 results in a heterochromatin structural change to the epigenome and the exclusion of RNA polymerase from the genome, preventing gene expression.
19Methyl groups derived from one-carbon metabolism are donated by a S-adenosylmethionine (SAM) dependant histone methyltransferase to a specific histone residue.
19 Dietary consumption of high folate foods, fortification or supplementation with cofactors of one-carbon metabolism such as folate and B12 contribute directly to histone methylation.
2,55Histone demethylation
Histone demethylase enzymes erase the previously written methylation of the histone residue induced by the histone methyltransferase.
19The enzymatic demethylase reaction by Jumonji C (JmjC)-domain-containing demethylases produce succinate and carbon dioxide, and is dependent on the presence of oxygen (O2), Fe(II) and the endogenous tricarboxylic acid (TCA) metabolite alpha ketoglutarate (aKG).
56 Dietary derived ascorbate was also revealed to be an essential cofactor for JmjC-domain-containing histone demethylases
57 which is supported in stem cell culture, with vitamin C treatment reducing global H3K9me2.
58 Demethylase reactions are showing to be necessary for natural killer cell activation and expression of interferon-y (INF-y) in the anti-viral response.
59 Inhibition of demethylase results in upregulation of glycolytic genes and downregulation of proinflammatory cytokines.
59 Hypoxia and environmental substances such as nitric oxide (NO) have also demonstrated down-regulation of histone demethylase activity.
29Mitochondria and Epigenetics
Beyond acetyl group synthesis, a cell’s energy production relies on acetyl-CoA entering the tricarboxylic acid cycle (TCA) for the production of adenosine triphosphate (ATP) and a variety of metabolites that also regulate epigenetic mechanisms, such as succinate and akG.
139 Mitochondrial oxidative phosphorylation is the primary source of cellular ATP.
160 Many epigenetic post-translational modifications depend on ATP, including chromatin remodeling complexes,
161 ubiquitination,
162 and phosphorylation.
163-166MAT is a well-documented ATP dependant enzyme.
164 This enzyme catalyzes the last step in the formation of SAM from methionine for both histone and DNA methylation epigenetic modifications.
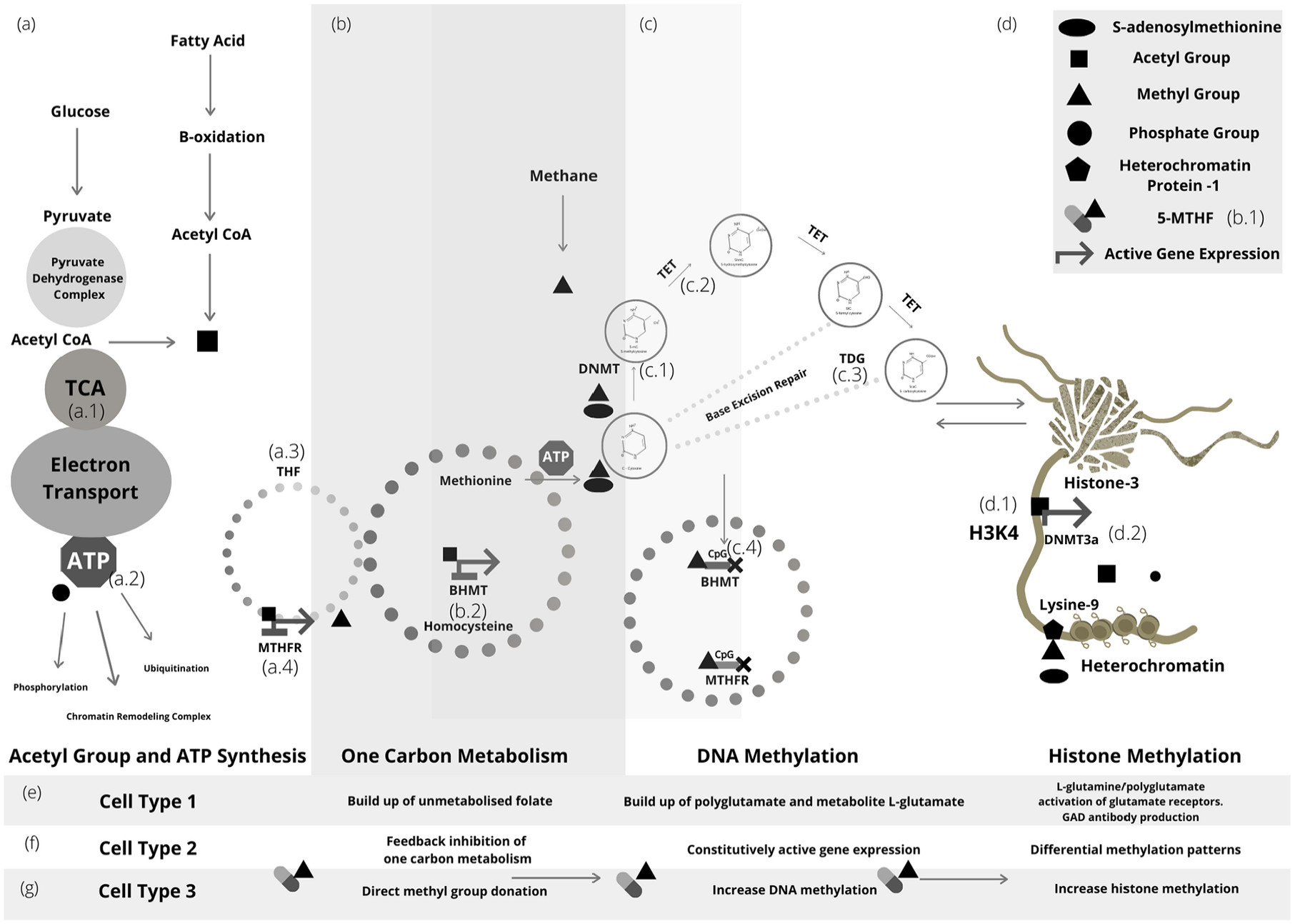
164 Figure 2 depicts some of the many cross-talk combinations that exist between primary epigenetic modifiers.
165-170For example, unmethylated histone-3 lysine-4 (H3K4) acts as an allosteric activator of DNA methyltransferase (DNMT3a) activating DNA methylation making H3K4 an auto-regulator of de novo methylation, and similarly, DNA methylation or histone methylation of genes within the one-carbon metabolic pathway reduces methyl group transfer and ultimately DNA or histone methylation.
Energy metabolism
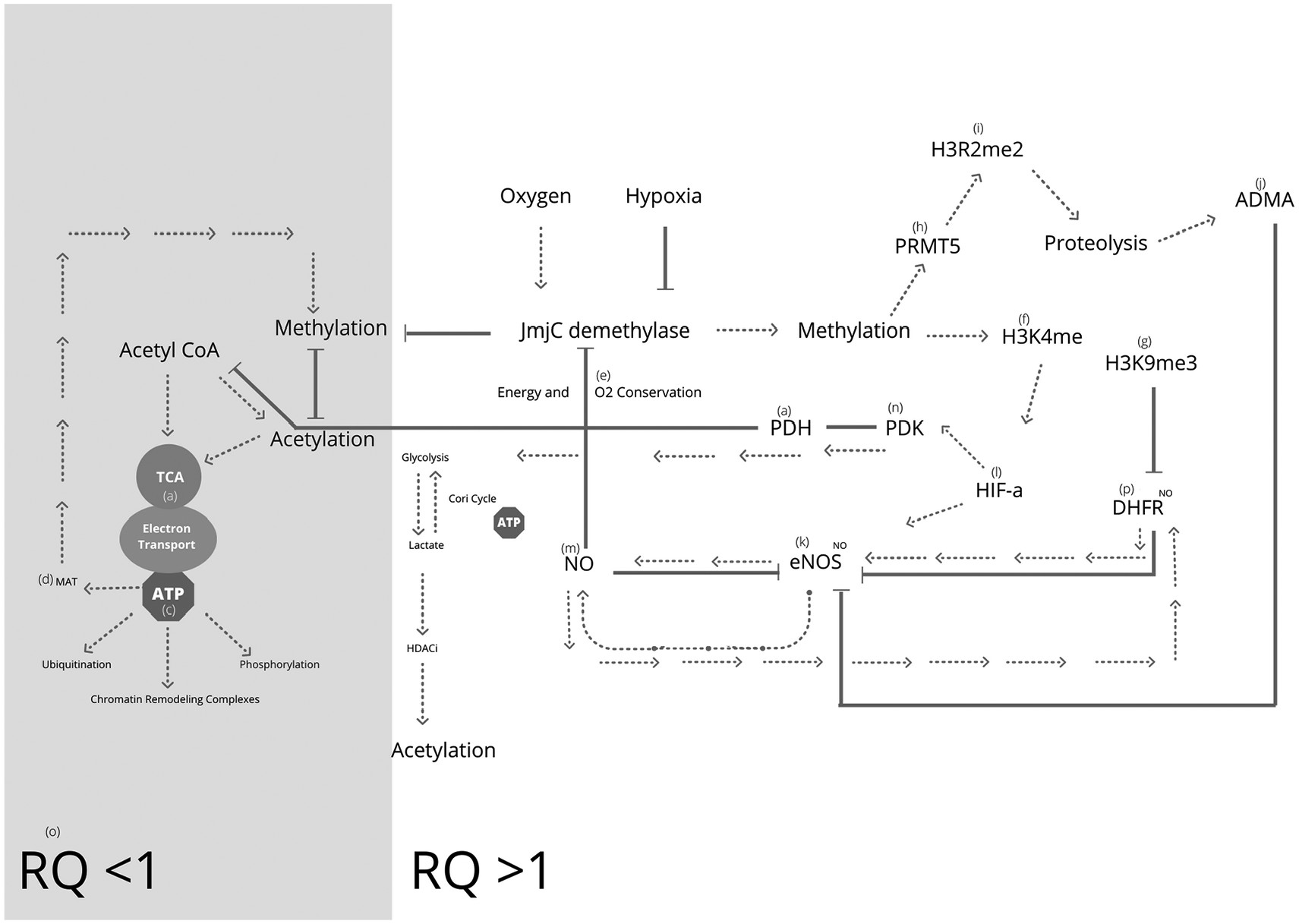
Energy metabolism can also be used to demonstrate epigenetic cross-talk (
Figure 3). For example; respiratory quotients (VCO2/VO2) have been used to estimate the use of energy substrate during rest and exercise.
171 RQ > 0.7 is said to represent mix substrate utilization, 0.8-0.9 protein utilization, 1 glycolysis only and >1.2 anaerobic metabolism.
172 Making RQ below 1 desired for high energy output and sustainability.
JmjC histone demethylase enzymes have demonstrated direct cellular O2 sensing.
173 A reduction in histone de-methylase substrates akG or O2
56 results in potent inhibition of histone demethylation and upregulation of repressive H3K9me2,
174 and activating H3K4me
174 resulting in reduced expression of DHFR
174 and upregulation of hypoxia-inducible factor a (HIF1a) respectively.
174 Upregulation of HIF1a is well known for its oxygen conservation via feedback inhibition of pyruvate dehydrogenase (PDH) activity, through upregulation of pyruvate dehydrogenase kinase (PDK) resulting in increased lactate synthesis and a reduction in the available acetyl CoA to enter the TCA and mitochondria for sustainable ATP synthesis,
175 subsequently increasing RQ to 1 or >1.2, resulting in excessive muscle lactate. Moreover; upregulation of HIF1a has been associated with the upregulation of eNOS,
176,177 resulting in increased NO and greater downregulation of demethylase activity
29 and O2 conservation.
B vitamins have long been associated with energy production; however, the mechanisms have remained undefined. The above describes how a reduction in sustainable ATP synthesis, through hypoxia, or hypermethylation at the level of acetyl CoA synthesis could result in downregulation of all primary ATP dependant epigenetic modifiers, relying on environmental or supplemental direct methyl group donation to drive methylation and glycolysis only metabolism (RQ > 1) which is unlikely to be sustainable for exercise such as endurance. Moreover; TCA cycle or mitochondrial ATP synthesis defects have been repeatedly implicated in diseases of cancer and dysfunctional immunity.
178As a result the excessive lactate production and consequent HDACi
153 may result in acetylation of undesired genes as seen in the metabolic switch known as the Warburg effect, which is commonly detected in cancer cells.
179Chronic fatigue syndrome (CFS)/Myalgic encephalomyelitis (ME)
A reduced ability to produce ATP, which is often characterized as deregulation of glycolysis at the pyruvate dehydrogenase complex is seen in patients with post-exertional malaise, or CFS.
16 McGregor and colleagues revealed changes in glycolysis and concentrations of metabolic pyruvate, acetate and lactate influencing both acetylation and deacetylation which were indicative of hypoacetylation in patients with CFS/ME.
16Epigenetic fluctuation
Immune cells fluctuate levels of epigenetic modifiers.
180 Specifically, T cells require coordinated suppression of methylation for memory T cell differentiation and appropriate response to antigen.
180 Similarly, the menstrual cycle
181 and circadian rhythms
182 require coordinated and a timely fluctuation of epigenetic modifications for regularity and synchronicity. Therefore, uncoordinated supplementation of methyl donors or modern epigenetic modifiers may result in dysregulated menstrual cycles, abnormal sleep patterns or immune dysfunction.
Nitrosylation
Tetrahydrobiopterin (BH4) content is maintained by the salvage and regeneration pathways.
186 Salvage of BH4 by dihydrofolate reductase (DHFR) promotes endothelial-derived nitric oxide synthase (eNOS) coupling and ultimately NO synthesis.
187 Like methylation; histone nitrosylation is demonstrating to be self-regulating, where the nitrosylation of eNOS inhibits its own expression,
188 and the nitrosylation of DHFR stabilises the protein and prevents it from ubiquitination and degradation increasing NO synthesis (
Figure 3).
189 BH4 deficiency or inhibition of DHFR by the methylated folate analog methotrexate (MTX) has demonstrated eNOS uncoupling resulting in the production of superoxide anions.
189 Similar to the hypoxic lack of O2 substrate and downregulation of demethylation; NO has also demonstrated direct inhibition of jmjC demethylase activity.
29 Interestingly; this may explain the downregulation of DHFR during hypoxia by providing direct superoxide production which has also demonstrated epigenetic modulation
190 and indicates a highly precise epigenetic regulation of gene expression, protein availability and energy conservation in the hypoxic state (
Figure 3).
Like the environmental methyl radicals, exogenous nitrogenous radicals may also influence epigenetic nitrosylation and ultimately the evolution of NOS.
191Animal-derived peptides such as those extracted from whey protein also play a role in epigenetic nitrosylation due to the up-regulation of nitric oxide synthase.
192Endothelial damage
Two primary mechanisms that have been implicated in the development of endothelial dysfunction are elevated levels of asymmetric dimethylarginine (ADMA) and a lack of dimethylarginine dimethylaminohydrolase (DDAH) enzyme activity.
193 ADMA is an endogenous eNOS inhibitor derived from the proteolysis of methylated arginine residues following arginine methylation by a group of epigenetic modifying enzymes referred to as protein-arginine methyltransferases (PRMT).
193 PRMTs are dependent on the one-carbon metabolite SAM for methyl group donation.
194 Increased ADMA has demonstrated superoxide release, uncoupling and inhibition of eNOS activity by up to 40%, and its presence predicts cardiovascular mortality, endothelial dysfunction in hypertension, hyperlipidemia, diabetes, and coronary artery disease.
193In contrast to MTX, in vivo vascular infusion of 5-MTHF has demonstrated improved endothelial function, eNOS coupling and decreased superoxide production.
195 However; in other cells, overexpression of eNOS has exhibited both male and female infertility.
196,197 Validating the unpredictable nature of orally administered 5-MTHF.
Phosphorylation
Inorganic phosphate for histone and protein phosphorylation are stored in the form of phosphocreatine, and ATP.
163 Phosphorylation histone -3 tyrosine-41 (H3Y41) by members of the Janus Kinase (JAK) family introduces a negative charge at the histone, excluding heterochromatin protein 1a (HP1a) from binding to H3K9me3
200 resulting in euchromatin and gene accessibility. Janus Kinase-2 (JAK2) gene expression is essential for hematopoietic differentiation
201 and many biological processes, most notably within the immune system.
202 Hypermethylation or loss of function of various genes within the JAK/STAT pathway have been implicated in a range of immunological conditions including recurrent staphylococci infection and allergic disease.
203-205 For example; DNA hypermethylation of JAK2 is responsible for dampened host immune responses in patients with tuberculosis.
205Protein phosphorylation of the primary immune transcription factor T box protein (T-bet) is essential for T helper cell 1 (TH1) differentiation.
206 Knockdown of phosphorylation or T-bet results in impaired T helper cell 2 (TH2) suppression resulting in the allergic phenotype.
207,208 T-bet induction of interferon-gamma (IFN-y) for viral immune response is dependant on H3K9 acetylation at the IFN-y locus,
208,209 which would not be possible without JAK2 exclusion of HP1a and prevention of H3K9me3.
200 Moreover; it has been determined that the histone lysine N methyltransferase (SUV39H1) dependant H3K9me3 incorporation of HP1a and the subsequent heterochromatin transcriptional silencing maintains the stability of TH2 cells and the induction of T-bet results in a resolution of allergic inflammatory lung pathology.
209Myeloproliferative neoplasms
An acquired JAK2 V617f mutation is present in the majority of patients with myeloproliferative neoplasms.
209 Several phenotypical manifestations result from the same mutation, and the presence of this mutation does not always present with disease.
210 The phenotypical presentation is dependent on the JAK2 burden, which is the accumulative JAK2 RNA from JAK2 copy numbers and the ratio of JAK2 RNA between wild type and its mutants.
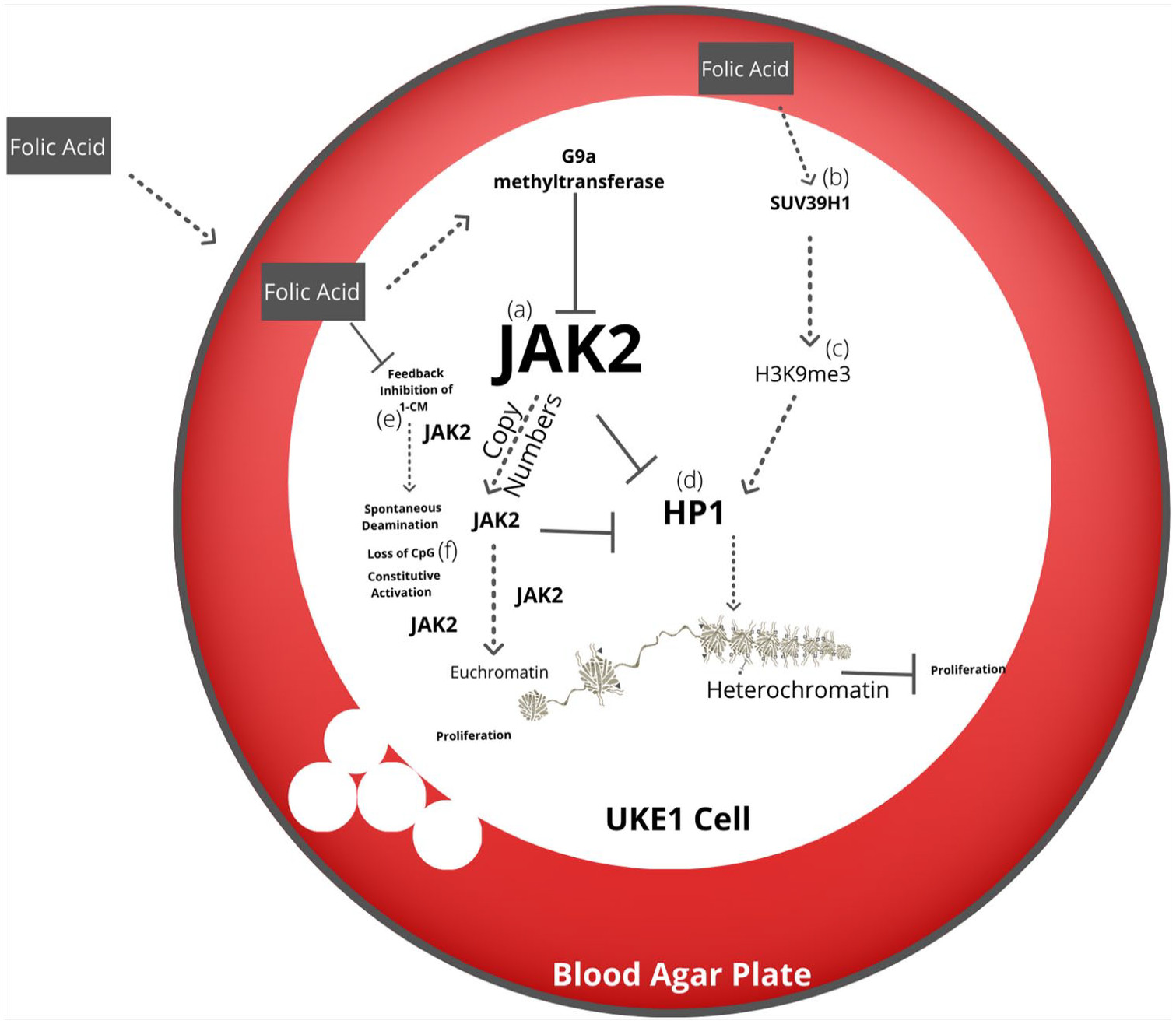
211 However, the neoplastic cell line UKE-1 shows the accumulation of JAK2 copy numbers in culture, suggesting epigenetic adaptation to the media.
209As previously described, human fibroblast cell lines frequently deaminate, losing methylation marks resulting in longevity or immortalization of cells.
76 Negative regulation of JAK2 is demonstrated by the upregulation of G9a methyltransferase at H3K9me2, and comparably the inhibition of G9a H3K9 methyltransferase increases JAK2 expression and ultimately H3Y41 phosphorylation, demonstrating direct epigenetic modulation by methylation.
201 Similarly, in cell culture, folic acid and the folate analog methotrexate also inhibit JAK2 phosphorylation.
212,213Like other cell lines, nutritional media for the UKE-1 cell line is also abundant in epigenetic modulators such as folic acid capable of supporting epigenetic inhibition of JAK2 and potentially a feedback loss of JAK2 methylation.
210,214 This suggests that chronic epigenetic modification may influence genomic adaptation, resulting in increased gene copy number or lead to constitutive activation of oncogenes such as JAK2 (
Figure 4).
Diet and Peptides
Many minerals, culinary, botanical herb, and spice phytochemical constituents, such as curcumin, resveratrol, quercetin, and sulforaphane have demonstrated powerful epigenetic modification in cell culture.
32,215 The fat-soluble vitamin retinol (vitamin A) has been described to drive epigenetic DNA methylation erasure through potentiation of TET demethylation enzyme activity.
216Vitamin D has also demonstrated DNA demethylation regulation through upregulation of jmjC domain-containing demethylase activity in cell culture; however, the cell-specific mechanisms are yet to be elucidated.
217,218Vitamin K has also demonstrated epigenetic modification in cell culture, producing histone deacetylase inhibition and hyperacetylation for treatment of cultured cancer cells.
219,220Animal-derived peptides have also demonstrated modulation of the epigenome with disease treatment potential in the laboratory.
221 Animals have very similar genomes to humans, and therefore animal-derived peptides from specific organs, sex-specific animals, animals with high metabolic rates or cuts of meat that produce specific proteins may increase the specificity of nutritional epigenetic modulation for treatment of disease.
Future Directions
Nutritional deficiency
Reduced erythrocyte, serum or plasma nutrient levels comparable to the average cohort have long been used to identify a nutritional deficiency. However, it is rarely considered that a reduction in an epigenetic modifier such as folate may be due to excessive use of substrate and enzymatic activity—for example; folate depletion due to excessive DNA methylation. The methylation requirement for each cell is different, making an evaluation of serum folate difficult to predict the level of folate within and required for glial cells compared to cardiomyocytes. Therefore; nutritional epigenetic research must look at nutrition from both angles keeping bio-individuality in mind, and practitioners should use caution in interpretation.
Rhythmicity and cell specificity
Outlining the cell-specific epigenetic regulation of rhythmic and coordinated biological processes will add to precision and patient management accuracy.
Laboratory techniques
Nutritional modulation of the epigenome in the laboratory is becoming increasingly popular and has profound disease modulatory potential. The cell, tissue and dose-specific epigenetic mechanisms for many environmental substances including bioactive peptides are yet to be determined; however, when it comes to experimentation, researchers should pay special attention to detail with the use of excipients, extraction, storage, reagents, media, and temperature as to avoid unintentional epigenetic modification and human error.
Accurate genotyping with the use of next-generation sequencing technologies is still an expensive operation, and the use of first-generation micro-array genotyping technologies is affordable and provides valuable insight into a patient’s ancestry and origin for precise nutritional health management. However; the use of restriction enzymes in this technique makes it impossible to differentiate a deletion from a methylated base or unrepaired deaminated base. Moreover; this technology does not provide information pertaining to haplotype selective expression, cell-specificity or gene epigenetic regulation, all of which largely affect the pathogenic effect of an SNV. This makes nutritional epigenetic modulation of specific genes based on these results currently unreliable, and therefore standardization of accurate genotyping is of great importance. It is now understood that regardless of genotype, epigenetics is the foundation of a gene’s regulation and therefore, RNA based sequencing techniques may provide a more reliable insight into cell-specific gene expression.
Novel epigenetic post translational modifications
A deeper understanding of the novel epigenetic post-translational modifications such as serotonylation and dopaminylation will aid suicide prevention and the natural management of mental health conditions and addiction.
Evolution
Understanding the impact of environments on gene evolution and epigenetics within populations or cultural subgroups should take priority as this opens up bio-individuality moving away from nutritional conformity and greater health for all populations.
Conclusion
With advances in genomic sequencing technology and the evolution into personalized medicine, nutritional epigenetics is showing to be an attractive non-invasive natural alternative to gene editing for the treatment of disease.
Insufficient MTHFR enzymatic activity is said to account for insufficient DNA and histone methylation, whereas an excess of DNA methylation and its etiology has been less explored. This is likely due to the difficulty of monitoring the regulation of the epigenome in cell culture.
Depending on the cell’s bio-uniqueness, the addition of 5-MTHF to the media may result in feedback inhibition of one-carbon metabolism and a loss of methylation or progresses to DNA methylation resulting in repression of gene expression, two completely opposing results, making nutritional supplementation without a thorough understanding of the patient’s bio-individuality unreliable.
The rapid epigenetic fluctuation that drive core human biological processes requires coordinated activation and deactivation of all potential modulators, suggesting prolonged supplementation with methyl group cofactors or other supplemental epigenetic modifiers may result in epigenetic inflexibility and metabolic blockages, providing symptom relief and disease concomitantly, which may or may not be immediately recognized. Knowledge of such paradoxical mechanisms emphasizes the importance and the need for highly specific personalized patient management.
This conceptional research has demonstrated that the underappreciated acetylation and downstream epigenetic post-translational modifications may be the pinnacle of the epigenomic hierarchy and that many variables are possible when it comes to epigenetic regulation of gene expression and DNA methylation of any primary epigenetic modifier has the potential to cause disease. Implications of excessive cellular methylation has been unclearly defined; however, this research demonstrates that an excess of methylation in cell culture results in epigenetic adaptation, immortality, differential methylation patterns, advanced evolution and a loss of CpG. Environmental factors have influenced the increasing incidence of reduced function MTHFR alleles through natural selection and bypassing evolution’s natural slowing of methyl group synthesis with methylated B vitamins may result in dysregulated feedback inhibition of one-carbon metabolism with unpredictable consequences. Given that methylation is directly antagonistic to acetylation upon the histone suggests that an excess of cellular methylation can result in significant chromatin inaccessibility and chronic gene inactivation. Moreover; hypermethylation of acetyl CoA production or a reduction in mitochondrial ATP synthesis due to hypoxia may result in epigenetic inflexibility, leaving primarily supplemental or environmentally derived methyl groups to drive aerobic glycolysis resulting in inefficient ATP production, and an excess of lactate.
It has become overwhelmingly clear that our genomic complex adapts to environment and diet, and the evolutionary consequence of differences between populations shapes our genomic blueprints resulting in subtle differences in the human phenotype. Human populations that undergo rapid dietary or environmental change tend to suffer the most with disease compared to those who remain true to tradition, such as centenarians of blue zone communities. Current international nutritional guidelines do not consider ancestry, bio-individuality or epigenetics so we must consider whether nutritional conformity may be negatively influencing the rate of evolution and contributing to disease.
The fortification of milled grains with B vitamins may have been implemented premature as the consequence of such intervention on other cells throughout differentiation, and its impact on neurodevelopment has been previously undefined. This report demonstrates that excessive oral or intravenous supplementation with 5-MTHF is non-cell specific, unpredictable and therefore may result in differential methylation patterns in cells with already sufficient cofactors.
In conclusion; this paper has explored the potential for nutritional epigenetics and has found personalized nutritional epigenetic modulation of disease through dietary, environmental, and lifestyle intervention to be a safe and highly effective option and should be a first-line treatment approach for the management and prevention of disease. In contrast; this report has established a requirement for cautionary measures in the interpretation of genomic sequencing data and revealed the irregularity and unpredictability for prolonged supplementation with epigenetic modifiers and the dangers associated with it. Therefore, the administration of epigenetic modulators through fortification or supplementation should be administered with caution and consideration for cell specificity, epigenetics, ancestry, and environmental influence.